Synthesis of Deoxyribonucleotides
| Home | | Biochemistry |Chapter: Biochemistry : Nucleotide Metabolism
The nucleotides described thus far all contain ribose (ribonucleotides).
SYNTHESIS OF DEOXYRIBONUCLEOTIDES
The nucleotides
described thus far all contain ribose (ribonucleotides). The nucleotides
required for DNA synthesis, however, are 2I
-deoxyribonucleotides, which are produced from ribonucleoside diphosphates by
the enzyme ribonucleotide reductase during the S-phase of the cell cycle.
[Note: The same enzyme acts on pyrimidine ribonucleotides.]
A. Ribonucleotide reductase
Ribonucleotide
reductase (ribonucleoside diphosphate reductase) is composed of two
nonidentical dimeric subunits, R1 and R2, and is specific for the reduction of
purine nucleoside diphosphates (ADP and GDP) and pyrimidine nucleoside
diphosphates (CDP and UDP) to their deoxy forms (dADP, dGDP, dCDP, and dUDP).
The immediate donors of the hydrogen atoms needed for the reduction of the 2I -hydroxyl group are two sulfhydryl
groups on the enzyme itself, which, during the reaction, form a disulfide bond
(Figure 22.12).
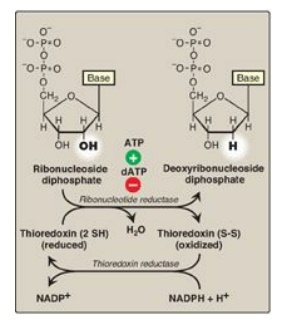
Figure 22.12 Conversion of
ribonucleotides to deoxyribonucleotides. NADP(H) = nicotinamide adenine
dinucleotide phosphate; dATP = deoxyadenosine triphosphate.
1. Regeneration of reduced enzyme: In order for ribonucleotide reductase
to continue to produce deoxyribonucleotides, the disulfide bond created during
the production of the 2 I-deoxy
carbon must be reduced. The source of the reducing equivalents for this purpose
is thioredoxin, a peptide coenzyme of ribonucleotide reductase. Thioredoxin
contains two cysteine residues separated by two amino acids in the peptide
chain. The two sulfhydryl groups of thioredoxin donate their hydrogen atoms to
ribonucleotide reductase, forming a disulfide bond in the process.
2. Regeneration of reduced thioredoxin: Thioredoxin must be converted back
to its reduced form in order to continue to perform its function. The necessary
reducing equivalents are provided by NADPH + H+, and the reaction is
catalyzed by thioredoxin reductase (see Figure 22.12).
B. Regulation of deoxyribonucleotide synthesis
Ribonucleotide
reductase is responsible for maintaining a balanced supply of the
deoxyribonucleotides required for DNA synthesis. To achieve this, the
regulation of the enzyme is complex. In addition to the catalytic (active)
site, there are allosteric sites on the enzyme involved in regulating its
activity (Figure 22.13).
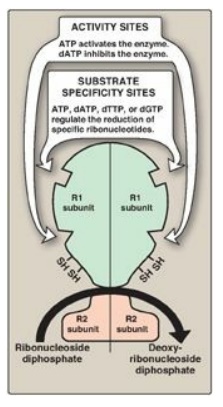
Figure 22.13 Regulation of ribonucleotide reductase. dATP = deoxyadenosine triphosphate; dTTP = deoxythymidine triphosphate; dGTP = deoxyguanosine triphosphate.
1. Activity sites: The binding of dATP to allosteric sites (known as the activity sites) on the enzyme inhibits the overall catalytic activity of the enzyme and, therefore, prevents the reduction of any of the four nucleoside diphosphates. This effectively prevents DNA synthesis and explains the toxicity of increased levels of dATP seen in conditions such as adenosine deaminase deficiency. In contrast, ATP bound to these sites activates the enzyme.
2. Substrate specificity sites: The binding of nucleoside
triphosphates to additional allosteric sites (known as the substrate
specificity sites) on the enzyme regulates substrate specificity, causing an
increase in the conversion of different species of ribonucleotides to
deoxyribonucleotides as they are required for DNA synthesis. For example,
deoxythymidine triphosphate binding at the specificity sites causes a
conformational change that allows reduction of GDP to dGDP at the catalytic
site.
The drug hydroxyurea (hydroxycarbamide) inhibits
ribonucleotide reductase, thereby inhibiting the generation of substrates for
DNA synthesis. Hydroxyurea is an antineoplastic agent and is used in the
treatment of cancers such as melanoma. Hydroxyurea is also used in the treatment
of sickle cell disease. However, the increase in fetal hemoglobin seen with
hydroxyurea is not due to its effect on ribonucleotide reductase.
Related Topics
