Resistance to Glycopeptide Antibiotics
| Home | | Pharmaceutical Microbiology | | Pharmaceutical Microbiology |Chapter: Pharmaceutical Microbiology : Bacterial Resistance To Antibiotics
Vancomycin and teicoplanin are the two glycopeptides used clinically. They bind the terminal D-alanyl-D-alanine side chains of peptidoglycan and prevent cross-linking in a number of Gram-positive organisms.
RESISTANCE TO GLYCOPEPTIDE ANTIBIOTICS
Vancomycin and teicoplanin are the two
glycopeptides used clinically. They bind the terminal D-alanyl-D-alanine side
chains of peptidoglycan and prevent cross-linking in a number of Gram-positive
organisms. They are not active against Gram-negative organisms because of the
presence of the outer membrane. Vancomycin use increased dramatically in
response to the increasing incidence of MRSA and resistance was first reported
in the enterococci in 1988. VRE now account for more than 20% of all
enterococcal infections. Resistance is greatest amongst Ent. faecium strains,
but significant numbers of the more clinically significant Ent. faecalis are
also resist-ant. Five types of resistance, VanA-VanE, have now been reported.
Phenotypic VanA resistance is the most common and confers high-level resistance
to vancomycin and teicoplanin. VanA resistance is mediated by a sevengene
cluster on the transposable genetic element Tn 1546(Figure 13.3).
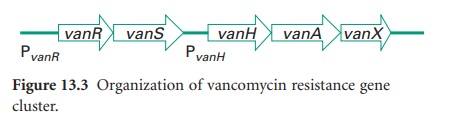
Resistance to vancomycin is via a
sensor histidine kinase (VanS) and a response regulator (VanR). VanH encodes a
D-lactate dehydrogenase/α-keto acid reductase and generates D-lactate, which is
the substrate for VanA, a D-Ala-D-Lac ligase. The result is cell wall precursors
terminating in D-Ala-D-Lac to which vancomycin binds with very low affinity.
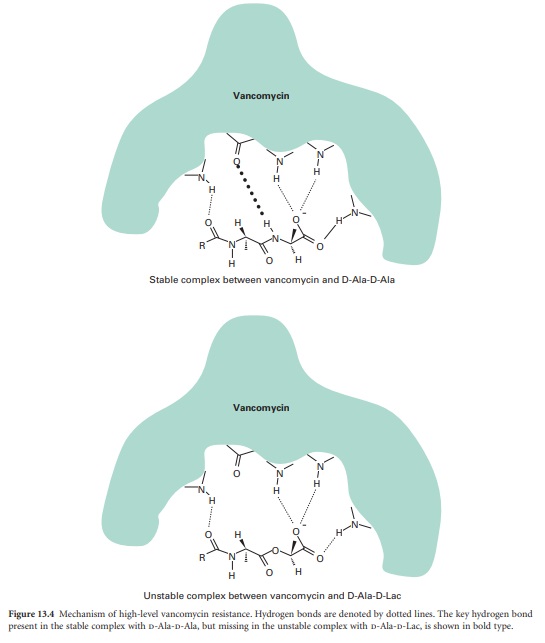
This change in affinity is mediated by one hydrogen bond. The complex formed
between vancomycin and D-Ala-D-Ala is stabilized by five hydrogen bonds,
whereas only four hydrogen bonds can form between vancomycin and D-Ala-D-Lac
and the complex is unstable (Figure 13.4).
Further, VanX encodes a D-Ala-D-Ala dipeptidase which can modify endogenous
D-Ala-D-Ala precursors. Recent genetic analysis has identified close homology
between this cluster and genes present in the vancomycin-producing
organism Amycolatopsis orientalis, suggesting that selective
pressure has forced genes originally present to protect antibioticproducing
organisms to jump to other species. VanB resistance is also acquired and the
peptidoglycan precursor is again D-Ala-D-Lac, but isolates often remain susceptible
to teicoplanin. VanC resistance is intrinsic and chromosomally encoded in some
enterococcal species such as Ent. gallinarum and the peptidoglycan precursor is
D-Ala-D-Ser. Less is known of VanD and VanE resistance, but both are acquired.
VanD uses D-Ala-D-Lac and VanE uses D-Ala-D-Ser.
MRSA And Reduced Glycopeptide Susceptibility
There is major concern that high-level,
VanA-type resistance could transfer to staphylococci, particularly MRSA.
Experimental transfer of the enterococcal VanA system to Staph. aureus on the
skin of mice has been reported, but other mechanisms resulting in
intermediate-level resistance occur in clinical isolates. In the 1960s and
1970s MRSA was not feared because several other treatment options existed,
including use of tetracyclines, macrolides and aminoglycosides. But multiple
resistance was accumulating and by the 1980s empirical therapy of
staphylococcal infections, particularly nosocomial sepsis, was changed to the
glycopeptide antibiotic vancomycin. MRSA levels were rising and the early 1990s
saw a major increase in vancomycin use. The inevitable consequence of the
selective pressure was the isolation in 1997 of the first Staph. aureus strain with
reduced susceptibility to vancomycin and teicoplanin (vancomycin MIC = 8μg/
ml). At the beginning of the 21st century, MRSA is responsible for up to 25% of
nosocomial infections in the USA and reports of community-acquired MRSA
infections are increasing. While reports of ‘superbugs’ resistant to all known
antibiotics abound, it is important to distinguish between reduced
susceptibility and resistance, recognizing that there are conflicting definitions
of resistance and resistance breakpoints. Strains with MIC values <4μg/ml
are considered sensitive, 8-16 μ g/ml intermediate and >32μg/ml resistant.
Thus the acronyms VISA (vancomycin-intermediate Staph. aureus) and GISA (glycopeptide-insensitive Staph. aureus) are used to
denote strains with vancomycin or teicoplanin MICs of 8μg/ml, whereas VRSA
(vancomyin-resistant Staph. aureus) is reserved for strains with MIC values
>32μg/ml. The mechanism of glycopeptide resistance is poorly understood, but
strains show longer doubling times and decreased susceptibility to lysostaphin.
Increased quantities of PBP2 and PBP2′
and cell wall precursors are presumed to trap vancomycin, while amidation of
glutamine residues in cell wall muropeptides reduces the cross-linking and
consequently the number of vancomycin target molecules.
Related Topics
